Genentech, a global biotech leader and member of the Roche Group, leveraged Snorkel Flow to extract critical information from lengthy clinical trial protocol (CTP) pdf documents. They built AI applications that used NER, entity linking, text extraction, and classification models to determine inclusion/ exclusion criteria and to analyze Schedules of Assessments. Genentech’s team achieved 95-99% model accuracy by using Snorkel Flow.
Unlocking the value of clinical trial protocol data
Scientists at Genentech and other life sciences companies write and perform thousands of Clinical Trial Protocols (CTPs) every year. These CTPs are complex documents that describe the plan for a clinical trial, including the objectives, the methodologies, and the population for the trial. There’s a lot of useful information in these CTPs that study design teams can reuse to reduce trial times and costs, increase recruitment of diverse patient populations and reduce the dropout rate of patients in a trial. If study teams have access to this data, their net outcome is a reduction in cost for drug development.

Genentech replaced months of manual data labeling using Snorkel Flow
Genentech started by using Snorkel Flow to build an AI application to extract 21 CMS Chronic Condition Entities from internal and external clinical trial protocols. Their application consisted of a named entity recognition NER model, an entity linking model, and a rules-based relationship extractor. Genentech leveraged programmatic labeling and a data-centric AI development approach to yield accurate inclusion-exclusion criteria that clinical scientists and study design teams used for analysis and data-informed protocol design.
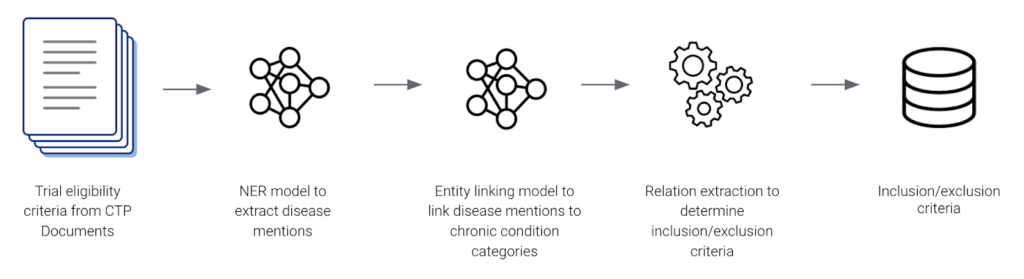
The data science team at Genentech built their end-to-end application pipeline in a few weeks achieving 98% accuracy with the help of Snorkel’s guided error analysis.
Snorkel Flow made the [clinical trial analytics] pipeline development adrag and drop experience. Michael DAndrea, Principal Data Scientist, Genentech
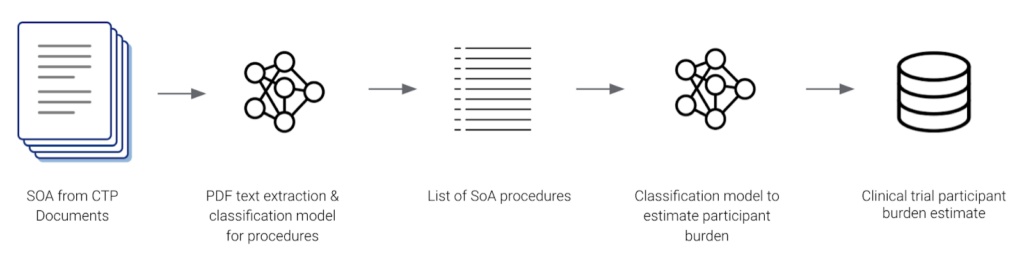
Genentech also used Snorkel Flow to build an AI application that estimated participant burden from CTPs. Their AI application identified and extracted procedure names from Schedule of Assessment tables and classified them into one of 8 categories. Since their data was labeled programmatically by Snorkel Flow, they were able to quickly adapt to changes in their label schema. The output data was used to harmonize terminology for clinical trial protocols across the organization.
By deploying AI applications built with Snorkel Flow Genentech estimates that they can increase recruitment for diverse populations and reduce clinical trial times costs and patient dropout rates. The combination of these outcomes will help Genentech dramatically reduce drug development costs and increase the number of drugs in their development pipeline leading to more cures.

This work was presented at the Future of Data-centric AI event hosted by Snorkel AI. Dive deeper into how Genentech used Snorkel Flow to build clinical trial analysis pipelines in this article.
