New Snorkel benchmark leaderboards. See the results.
Clinical entity classification in electronic health records
Research recap: Ontology-driven weak supervision for clinical entity classification in electronic health records (EHRs)
In this post, I have summarized the research published in this academic paper, Ontology-driven weak supervision for clinical entity classification in electronic health records by Jason Fries et al. This paper was published in Nature Communications in 2021.Problem statement
Electronic health records (EHR) contain a rich set of information such as clinical notes, laboratory results, diagnoses, among other things, that can be utilized to tailor a specific treatment for each patient. The extraction of entities such as drugs and disorders is an important step in making clinical decisions. Recently, natural language processing (NLP) techniques such as named entity recognition (NER) have been used for automating tasks such as identifying disease names or other entities from text.

Challenges with NER
Traditionally, training classifiers for named entity recognition (NER) and cue-based entity classification have relied on hand-labeled training data. However, annotating EHR requires considerable domain expertise and money, creating barriers to using machine learning. Moreover, hand-labeled datasets are static artifacts that are expensive to change. Due to privacy concerns regarding patient data, outsourcing and sharing labeled data is often a non-starter. Later in this research, the authors discuss the need for agile and robust techniques for acquiring training data for machine learning models in light of the fast-changing events of the COVID-19 pandemic.
Proposed method
The authors propose Trove, a framework for weakly supervised entity classification using medical ontologies and expert-generated rules. With this approach, instead of manually labeling training data, the authors use Snorkel’s weak supervision framework to programmatically label EHR. This makes it easy to share and modify training data, while offering performance comparable to learning from manually labeled training data.Weak supervision is the technique of generating low-cost and less accurate labels (called labeling functions) by utilizing subject matter expert heuristics, rules-based systems, large language models, dictionaries, and ontologies as multiple imperfect resources for supervision. Weak supervision has demonstrated success across a range of NLP and other settings at Google, Genentech, Intel, Apple, Stanford Medicine, and more.In this paper, Trove applies weak supervision by creating labeling functions using:
- Task-specific rules: extraction of rules and heuristics based on specific text such as physician notes
- Ontologies: extraction of terminologies from external resources such as Unified Medical Language System (UMLS 1), or other dictionaries
Trove pipeline
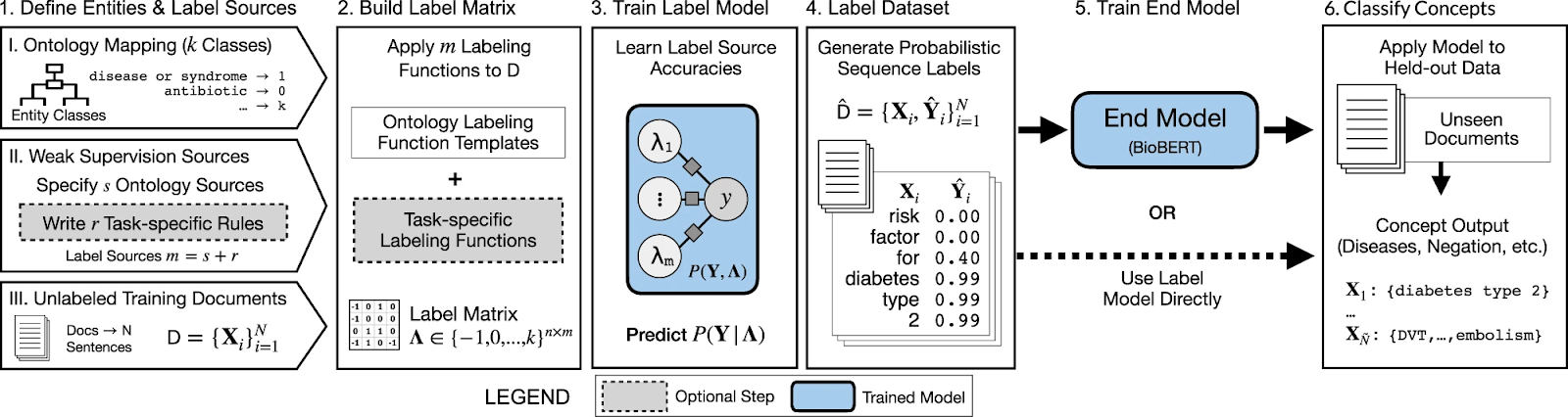
The following is a quick description of the Trove pipeline:
- User start by
- Specifying ontology dictionaries (UMLS, etc.), and then a mapping from ontologies’ categories to class entities labeling via ontologies and task-specific rules
- Collecting a large set of unlabelled training documents
- The label matrix is built by aggregating all labeling functions
- The label model is trained to correct the noise and learn the accuracies in labeling functions
- It predicts a consensus probability per word to generate a labeled dataset
- The labeled dataset is then used to train an end model such as BioBERT 2
- Or the label model predictions could be used as the final classifier
- Finally, apply the end model to process documents and obtain predictions per word
Datasets and tasks
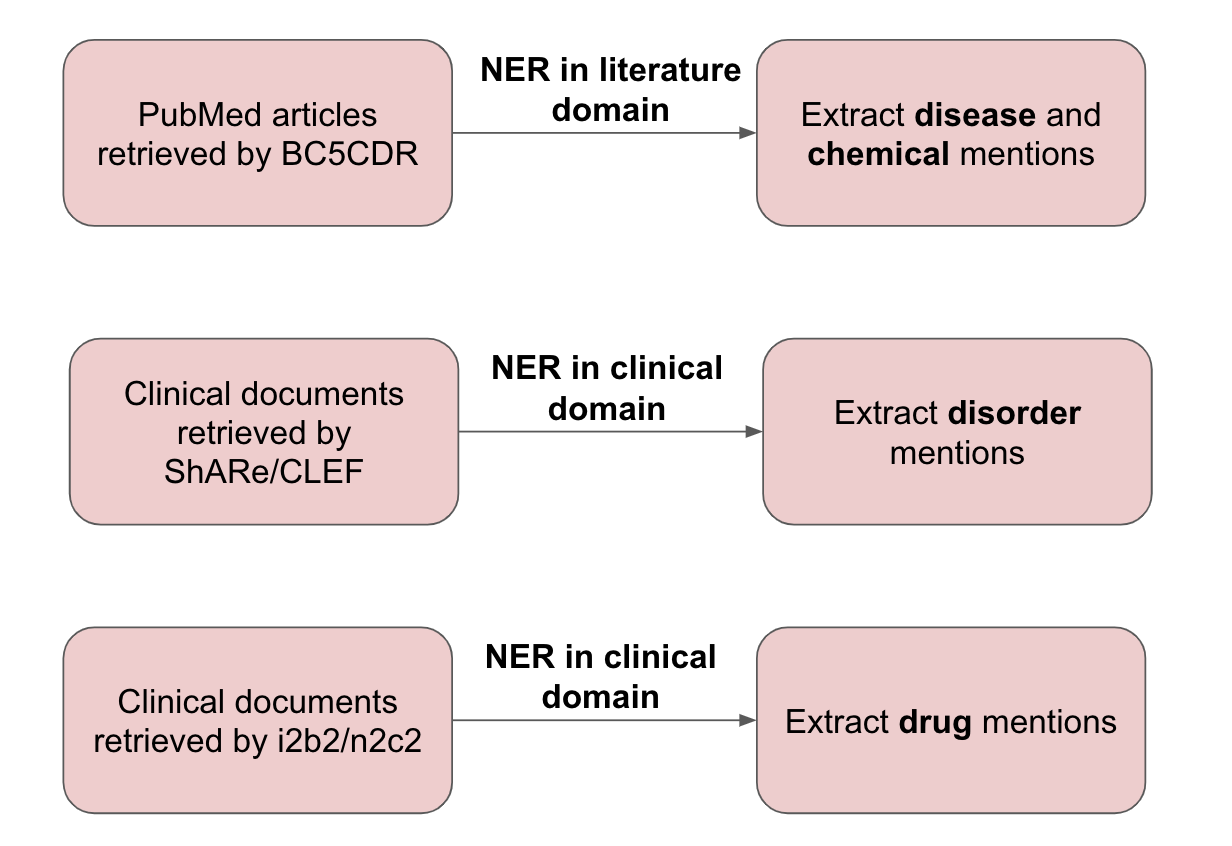
Several NER datasets are used in this study for chemical/disease and drug tagging.
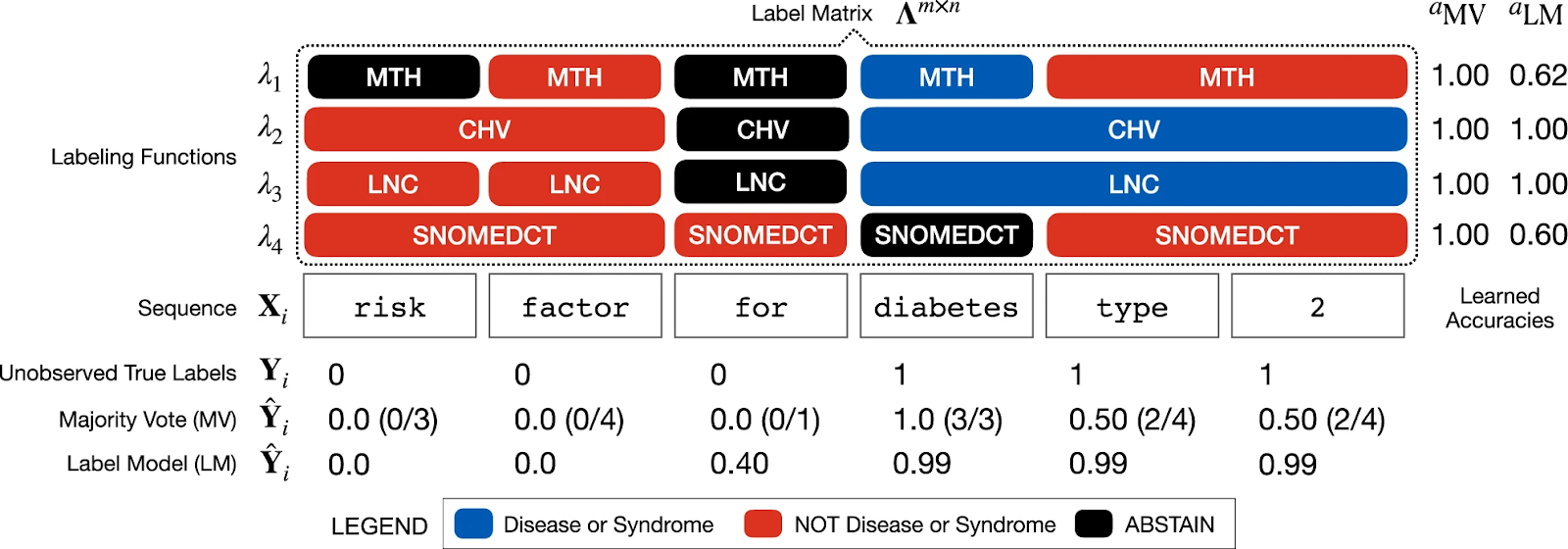
In the figure below, an example sequence X i is shown with the application of four different ontology labeling functions (MTH, CHV, LNC, SNOMEDCT). The entity of interest to be tagged is ‘diabetes type 2’.
- Majority vote estimates Y i as a word-level sum of positive class labels, weighing each equally
- Label model estimates Y i by reweighting labels to generate a more accurate prediction
Combining labeling functions and training a classifier; what’s the effect on performance?
To test Trove’s capabilities, several experiments have been performed using the datasets mentioned above. In the first experiment, the labeling functions are aggregated by majority vote, and the label model, and then the extra effect of training another classifier by weak supervision is explored. These performances are compared against fully-supervised models by having access to hand-labeled data for training, and published state-of-the-art performances. In summary, the following four methods are tested and compared on all four sequence tagging tasks:
- Majority vote (MV): the majority class vote for each word
- Label model (LM): the default output of data programming model
- Weakly supervised (WS): BioBERT trained on the probabilistic dataset generated by the label model
- Fully supervised (FS): BioBERT trained on the original expert-labeled training set
- State-of-the-art (SOTA): published performance metrics in the literature
Ontology-based vs. ontology-based AND task-specific labeling functions; what’s the effect on performance?
In the next experiment, the effects of different labeling functions are analyzed. Medical ontologies are a great source of information for weak supervision, however, additional task-specific rules may be required for a boost in performance. To test this, the following two approaches are explored for all four tasks:
- Using ontologies for labeling functions
- Dictionary of numbers, stopwords, punctuation
- + UMLS
- + Existing ontologies not in UMLS
- Using ontologies AND task-specific rules
- Regular expressions
- small dictionaries (e.g., illegal drugs)
- other heuristics
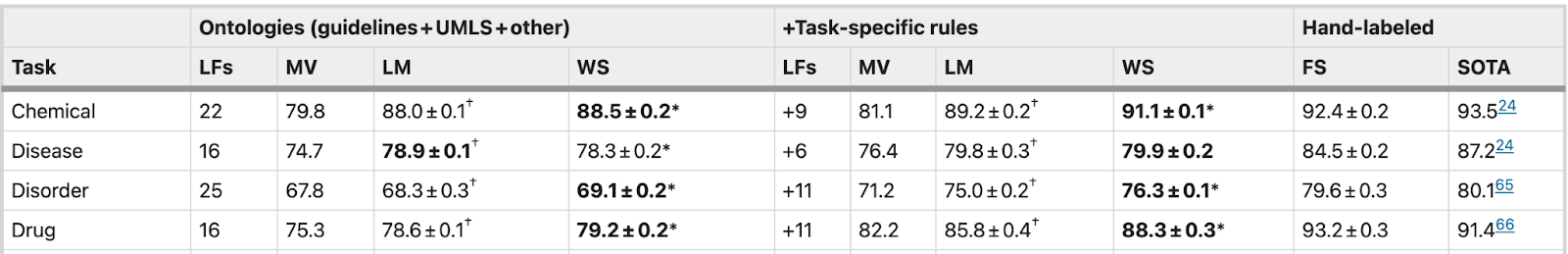
The table below summarizes the results of the application of different approaches:
- LM performance was higher than MV in all tasks by 4.1 F1 points on average
- Weak supervision using BioBERT provided an additional average increase of 0.3 F1 points over LM
- Weak supervision using additional task-specific rules performed within 1.3–4.9 F1 points (4.1%) of models trained on hand-labeled data (FS)
Case study: COVID-19 risk factor monitoring
COVID-19 pandemic presented a situation where there was a critical need to rapidly analyze literature and unstructured EHR data to fully understand symptoms, outcomes, and risk factors at short notice. Several challenges arise in making rapid classification models, such as manual labeling costs and data privacy concerns. This final case study of Trove is for COVID-19 symptom tagging and risk factor monitoring using a daily data feed of Stanford Health Care (SHC) emergency department notes.
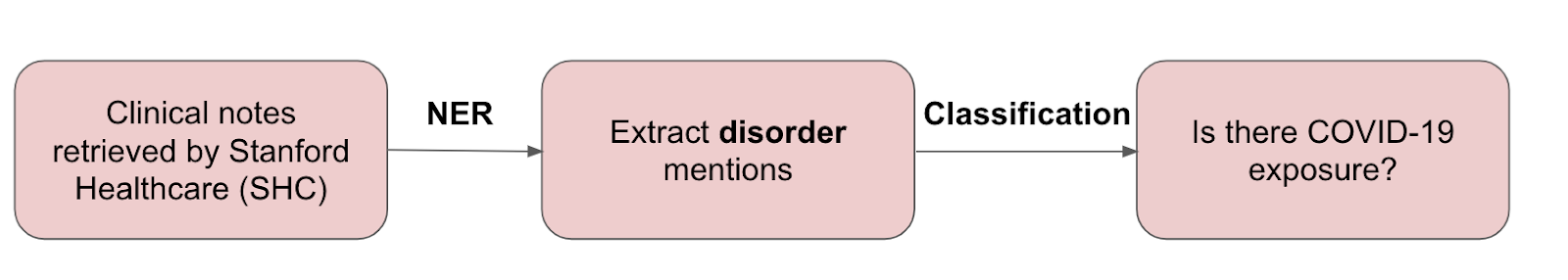
Results
Disorder tagging: Ontology-based weak supervision performed almost as well as a hand-labeled FS model while adding additional task-specific rules—outperformed the FS model by 2.3 F1 points.
Exposure classification: The weakly supervised end model provided a 5.2 F1 points improvement over the rules alone.
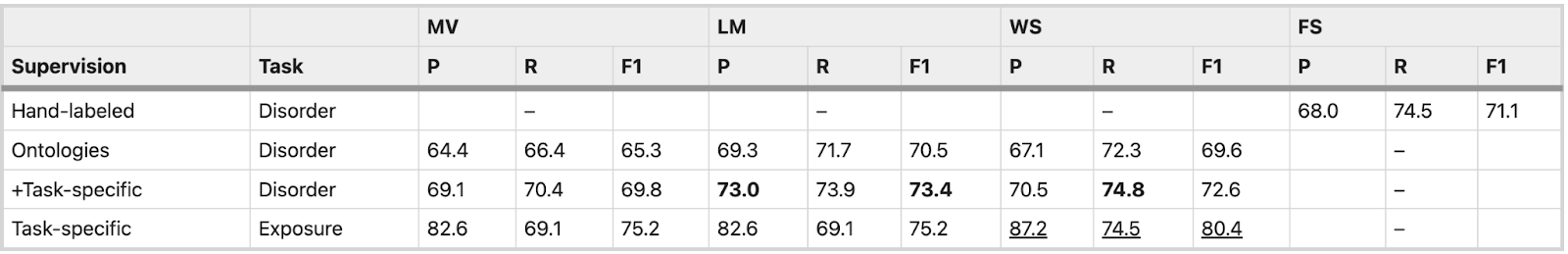
*P: Precision, R: Recall, F1: F1-score
Conclusion
The Trove framework demonstrates how classifiers for a wide range of medical NLP tasks can be quickly reconstructed.
- Fast: No need for time-consuming hand-labeled training data
- Explainable: Rule-based and ontology-based labeling functions provide an interpretable view of generated training labels
- Privacy preserved: Easy to share, edit, and modify labeling functions
- High performance: combining the state-of-the-art machine learning (such as BioBERT language model) with the flexibility of rule-based approaches offers performance comparable to learning from manually labeled training data
References
- Lee, J. et al. BioBERT: a pre-trained biomedical language representation model for biomedical text mining. Bioinformatics 36, 1234–1240 (2019).
- Bodenreider, O. The unified medical language system (UMLS): integrating biomedical terminology. Nucleic Acids Res. 32, D267–70 (2004).
Featured photo by Jeremy Bezanger.
Nazanin Makkinejad is an applied machine learning engineer at Snorkel AI, where she works with enterprise data science teams to realize the benefits of data-centric AI and Snorkel Flow. Prior to her role at Snorkel AI, Nazanin was a Postdoctoral Research Fellow at Harvard Medical School (HMS) and Massachusetts General Hospital (MGH), working on the intersection of deep learning and brain image analysis. She has a Ph.D. from the Illinois Institute of Technology in Biomedical & Medical Engineering and a Master’s Degree in Electrical and Computer Engineering from The University of Illinois Chicago.
