Doctors and medical professionals have long struggled to predict and understand the adverse drug events prescriptions can have on their patients, but machine learning and artificial intelligence can help doctors prescribe drugs more intelligently.
Adverse drug effects (ADE) can range from mild to severe, and can have significant impacts on patients’ health and well-being. Understanding the potential side effects of drugs is important for healthcare providers in selecting appropriate medication and reducing the risk of harm to the patient.
Physician notes, electronic health records, transcripts of calls between patients and their pharmacists, and other sources of unstructured data can offer insight into drugs’ negative side effects. However, mining this data manually is prohibitively expensive and time-consuming. Fortunately, modern machine learning tools (including named-entity recognition) can help automate information extraction, and Snorkel AI’s Snorkel Flow platform can help users build effective applications around these tools quickly.
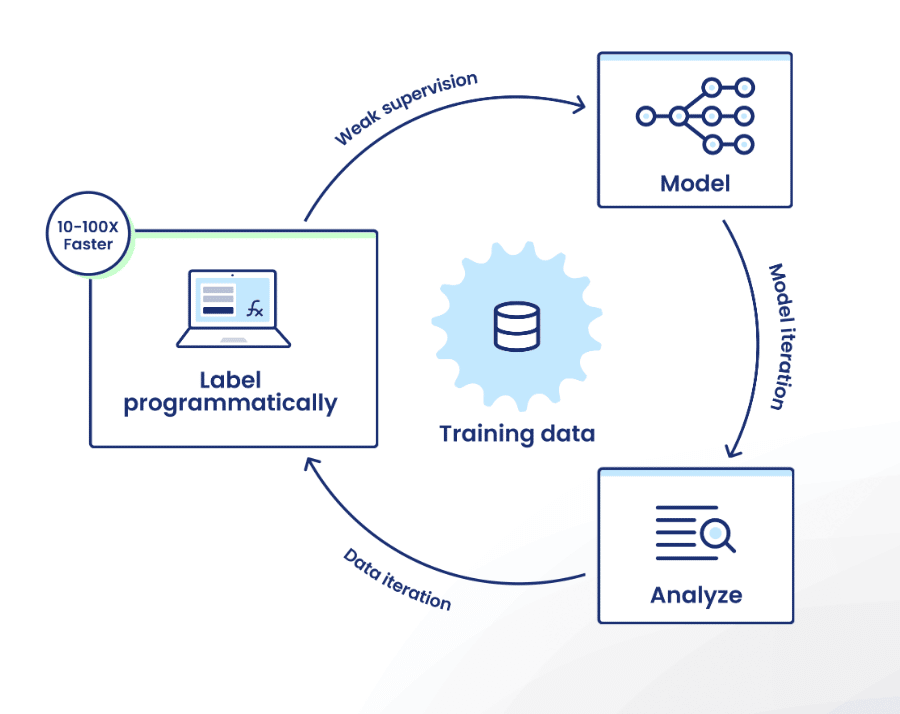
Platform capabilities
Snorkel Flow offers the ability to create high-quality training sets by encoding domain expertise as labeling functions. This lets users effectively label hundreds or thousands of data points at once—sometimes in less time than it would take them to label a single data point manually.
Users can then train a model using that data, and improve model performance by iterating on training data with directed error analysis.
Extracting drugs and ADEs using machine learning
In this task, we worked with two public datasets to identify named drugs as well as negative outcomes: CSIRO Adverse Drug Event Corpus (a corpus of medical forum posts), and ADE-Corpus-V2 (a corpus of PubMed abstracts), which in total included 1860 documents. An example input and output are provided below:
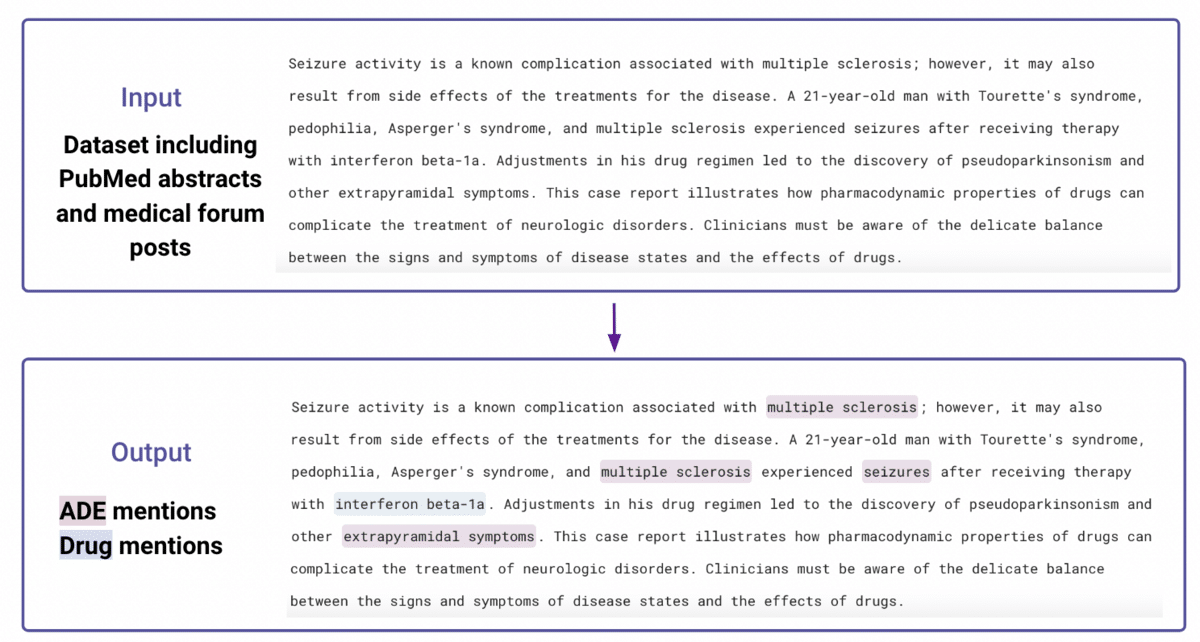

Step 1: Prepare a small amount of ground truth data
The dataset was split into train (80%), development (10%), and validation (10%) sets. Prior to starting our work, subject matter experts only labeled the development and validation sets to gauge the effectiveness of our labeling functions and unlock the full analysis workflow. The train set was fully unlabeled.
Step 2: Create baseline labeling functions using keywords
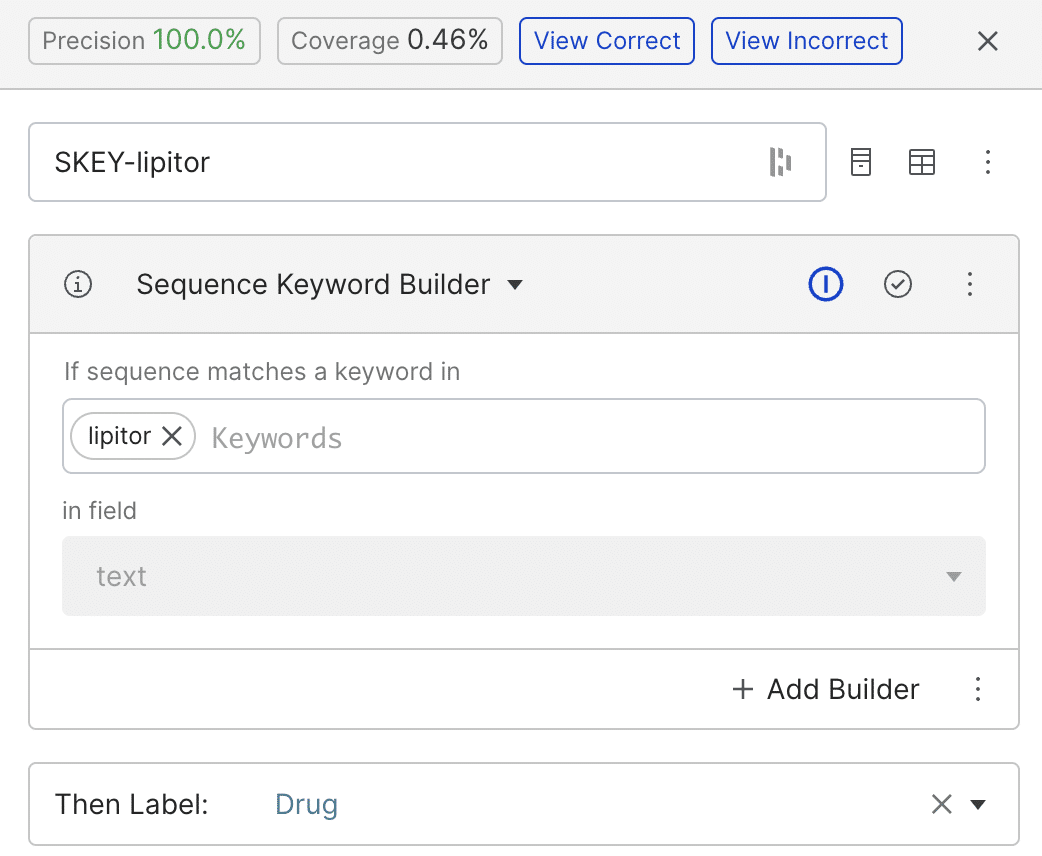
We started by building simple labeling functions (LFs) using patterns or regular expressions. In the example below, the keyword-based LF tags any span matching the word ‘lipitor’ as a drug. While the LF achieved a precision of 100%, its coverage is low.
We could solve our coverage shortfall by creating separate labeling functions for each drug and negative outcome we are interested in, but Snorkel Flow allows us to work more efficiently.
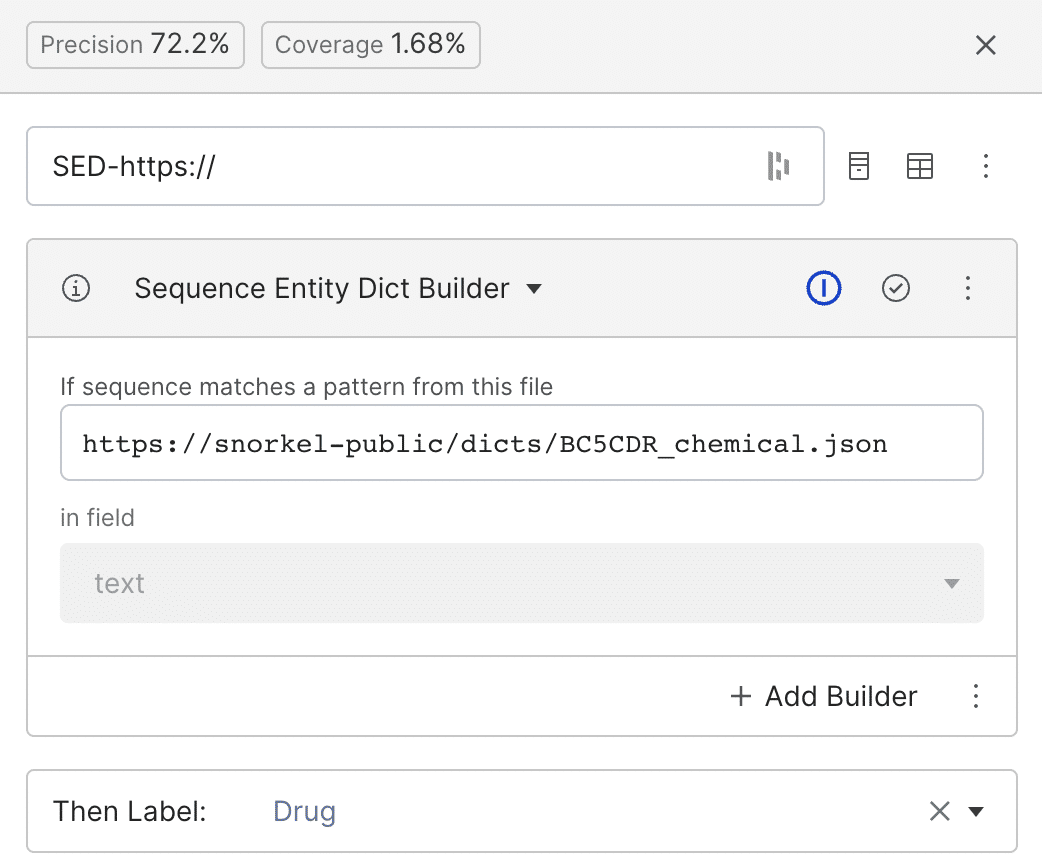
Step 3: Labeling functions that use dictionaries
To scale our labeling abilities faster, we can use public or private dictionaries that contain lists of keywords that frequently appear for each label. Here, we use a public dictionary based on BC5CDR (a dictionary of chemicals, diseases, and the interaction between chemicals and diseases). Every span in the text matching any of the “drug” entries in the dictionary will be labeled as class drug (Zetia, Lipitor, Naproxen, etc).
Step 4: First iteration to train a model and guided error analysis
Labeling functions do not need to have high precision or cover all the data. After writing a few LFs, a label model integrates them, denoises them, and assigns a probabilistic label to each data point in the training set. Then, we begin training our first preliminary models using the data set.
For this task, we used a base model derived from BERT (Bidirectional Encoder Representations from Transformers). BERT is a family of state-of-the-art tools for natural language processing and related tasks such as named entity recognition. Training a BERT model from scratch (or even fine-tuning BERT models) requires a large training dataset, which is not always available due to cost, privacy, and domain expertise, among other things. But we’ve just solved that problem using Snorkel Flow’s programmatic labeling.
We train our first model using DistilBERT (a smaller version of BERT) and see a low F1 score. We expected this; our keywords and dictionary labeling functions covered too little of the data to distill adequate amounts of signal from each class. Using guided error analysis and active learning, we identify areas of improvement and try to write more advanced LFs. Also, we realized that the ground truth is missing some obvious mentions of drugs and adverse events. For example, if the word ‘lipitor’ was mentioned twice in a document, only one occurrence may have been tagged. So, we tried to correct for such ground truth inconsistencies in the development and validation sets.
Step 5: LFs based on HuggingFace pre-trained models
We added more advanced labeling functions using the pre-trained HuggingFace models BioBERT and SciBERT. These models have been trained on medical publications and scientific papers, respectively, and come with useful off-the-shelf token-tagging abilities that we use as an ingredient in our labeling functions.
For example, SciBERT will tag certain tokens or words in a document as ‘DRUG.’ Hence, we can build a function that labels any token that SciBERT marked as ‘DRUG’ as a drug.
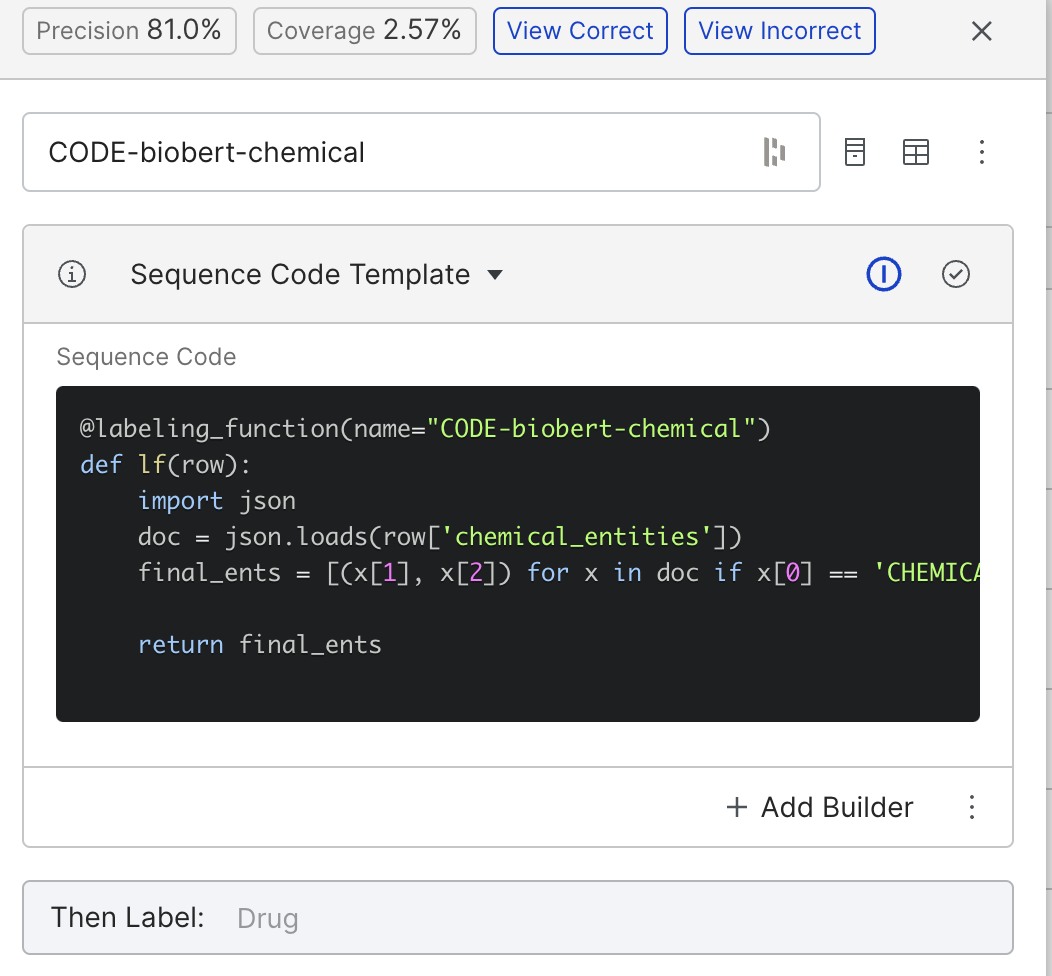
We can build something similar using BioBERT by labeling anything BioBERT tags as “CHEMICAL” as the drug class. The above function achieved a precision of 81%.
Step 6: LFs based on public ontologies
We were able to label additional entities using medical ontologies from the Wikidata API. Wikidata is a structured data representation maintained by the same organization as Wikipedia. When called, it returns a label, description, and a list of statements and identifiers for the passed string.
In order to use this capability, we first extracted noun chunks from each document using SpaCy and scispacy entities. We submitted those spans to Wikidata’s API, searched the returned list of identifiers for medical ontologies, and mapped them to the drug and ADE labels. Specifically, if ‘RXNorm ID’ or ‘DrugBank ID’ existed in the Wikidata search results, that span would be labeled as drug. Similarly, if we found the ‘Symptom Ontology ID’ code, we would label that span as an adverse drug effect.
We used the same strategy in obtaining ontology knowledge from the BioPortal API.
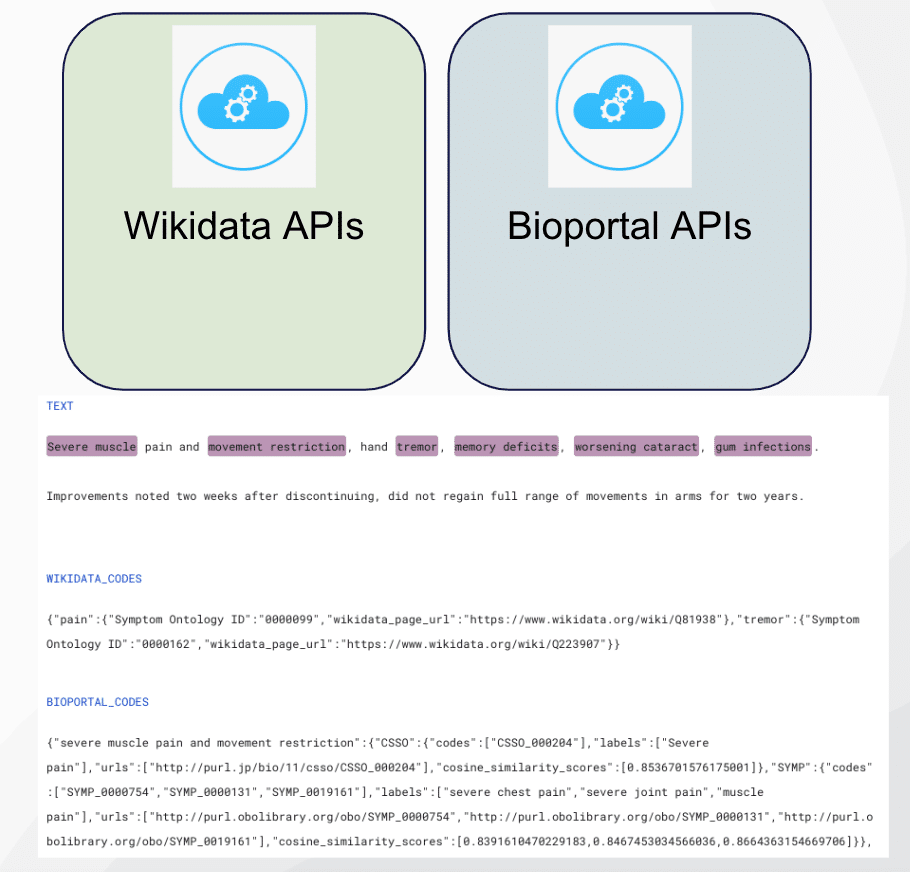

Step 7: LFs based on foundational or large language models
Large language or foundation models such as GPT-3, BERT, Stable Diffusion, and others have proved to be incredibly powerful tools for NLP tasks. Using Snorkel Flow, we can leverage any FM’s output as a source of signal in writing LFs.
For example, users can query foundation models with relevant prompts to extract domain knowledge efficiently. Here, we asked the deepset/roberta-large-squad2 model “What drug did the user take?” Using the confidence rate of the response, we’re able to auto-label drug examples. We could have used any other foundation model in this task.
LABEL = “Drug”
qa_prompter = QAPrompter(
node_info=NODE_INFO,
text_col=”text”,
label=LABEL,
)
loader = CachedLocalLoader(
llm_type=LLMType.QA,
hf_model_name=”deepset/roberta-large-squad2″,
use_gpu=True,
)
PROMPT = “What drug did the user take?”
qa_prompter.output_on_examples(
prompt=PROMPT4, loader=loader
)
Here is the output on an for labeling drugs.

Step 8: LFs based on cluster embeddings
Snorkel Flow lets us unlock the capability of embedding-based clusters as labeling functions. By having a small amount of ground truth, Snorkel Flow will automatically generate Cluster LFs that users can review and use as a source of signal.
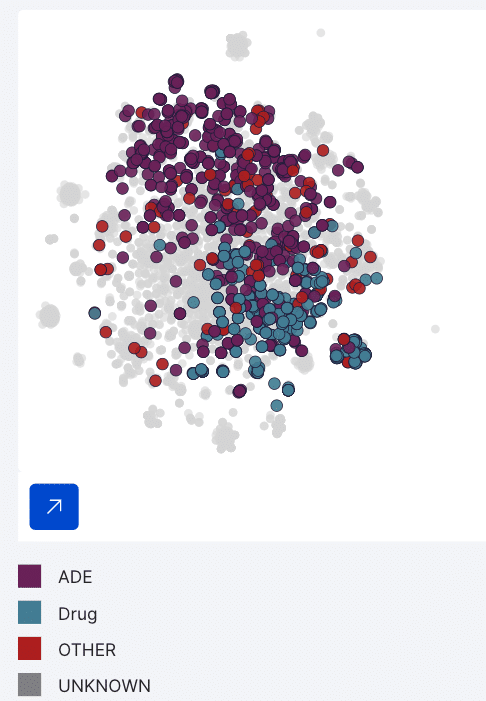
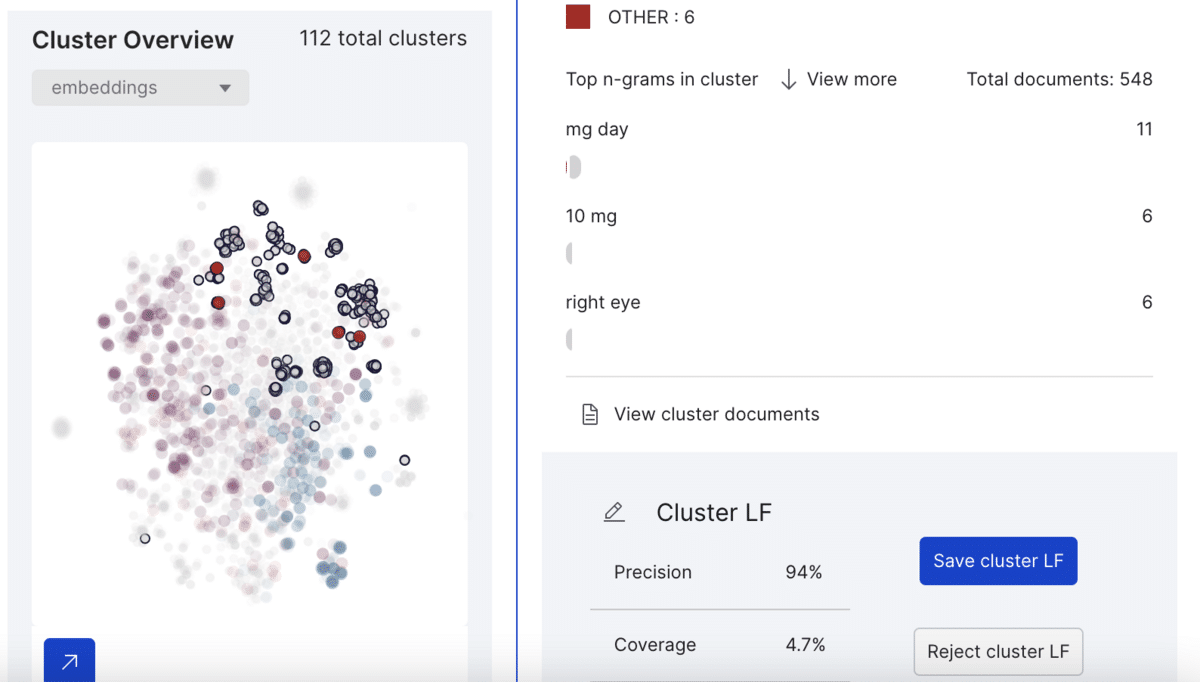
This example cluster contains 548 spans from which Snorkel Flow labels six as class other (neither drug nor ADE) with 96% precision and 4.7% coverage. Upon saving the cluster LF, the platform labels every other span belonging to this cluster as “other.”
Step 9: Re-training and drive to quality
After creating our new labeling functions, we used AutoML with DistilBERT to train a model using the data. We tested the performance on the manually-labeled validation set, and achieved an F1-score was 87% for the drug class and 71% for the adverse effect class—much higher than the initial rounds.
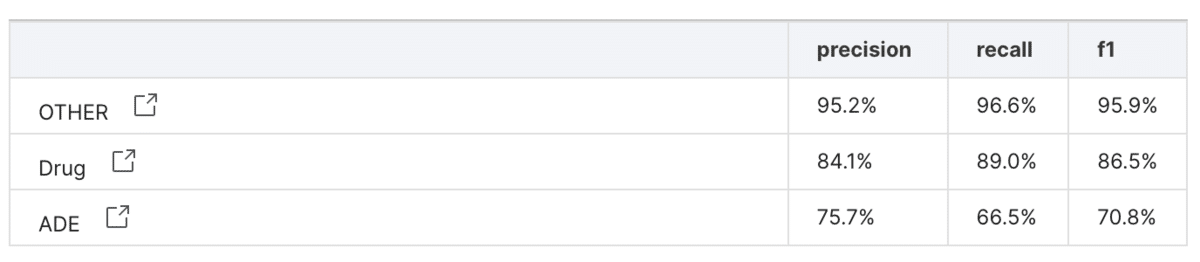
Comparison to manual labeling
To verify our achievements, another team performed this project without Snorkel Flow, asking subject matter experts to label the entire data set manually. Data scientists then trained a model on the manually labeled dataset and calculated its performance—which was 0.33 F1 on the class drug and 0.58 F1 on the class ADE. One important note here is that without Snorkel Flow, the data scientist and subject matter experts never realized there were inconsistencies or missing ground truth in their labeled documents.
The manual approach took about two weeks. Labeling with Snorkel took two hours, and that clear advantage scales with larger data sets.
This proof-of-concept used 1,600 unlabeled records. A real-world application that needed 100 times as much data would require subject matter experts to spend 200 weeks labeling. With Snorkel that time would still remain at 2 hours.
In addition, our rapid iterative loop and guided error analysis allow users to achieve high-performance models much faster.
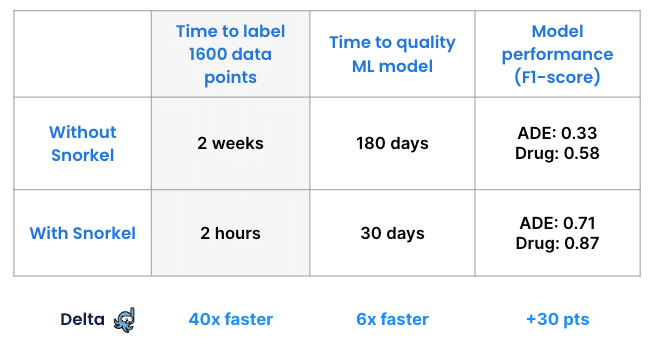

Conclusion
In this project, we used Snorkel Flow’s data-centric approach to build a named entity recognition model to identify drugs and adverse drug effects with high F1-scores. Without Snorkel Flow, getting access to a properly-labeled dataset for this task could be challenging or impossible. Instead, the platform allowed us to label data 40 times faster than manually labeling, and then quickly get to a high-quality DistillBERT model.
This model, enabled by this rapidly-labeled dataset, could enable the ability to analyze drugs and their adverse events at a large scale. Understanding this relationship could help healthcare providers to consider all potential side effects when they choose medications for their patients.
Also, by being able to build projects much faster and with lower costs, not only can healthcare providers shift their attention to more important tasks rather than manual labeling, but also leverage the high-performing models to make more informed decisions.
Learn More
Follow Snorkel AI on LinkedIn, Twitter, and YouTube to be the first to see new posts and videos!
